Thyroid nodules are common and frequently benign, with approximately 5–10% being malignant.1 Current diagnostic evaluation of thyroid nodules should include a multidisciplinary approach, integrating clinical, sonographic, cytologic, and molecular analysis. While fine-needle aspiration (FNA) biopsy and cytologic examination have emerged as the gold standard in separating benign from malignant nodules, cytology is limited and challenging when it is reported as indeterminate.2 Indeterminate cytology is reported in 10–20% of thyroid FNAs, with management options including observation, repetition of FNA, or diagnostic lobectomy.3 Most patients with inconclusive cytology are referred for a diagnostic lobectomy immediately or after a re-biopsy demonstrating persistently inconclusive cytology results, with cancer confirmed in only 20%.3,4 In this strategy, the majority of those undergoing surgical excision will have benign nodules, suggesting that this is not a cost-effective approach. To improve diagnostic and/or prognostic accuracy, molecular markers have been developed and employed to improve patient selection and reduce unnecessary surgery, while enhancing understanding of individualized prognosis.5
Enormous efforts in search of objective measures that can differentiate benign from malignant thyroid lesions have been recently undertaken, mainly involving immunohistochemical markers and molecular markers, using somatic mutation, gene expression, and microRNA (miRNA) analyses.6 Molecular analysis is a broad term, encompassing various types of molecular testing. Specific mutations arise from failure of DNA repair pathways, or in a repair-competent background resulted from genotoxic stress from cellular processes such as transcription and replication that overwhelm high-fidelity DNA repair. The patterns of multiple expressed genes that correspond to benign or malignant processes are identified by messenger RNA (mRNA) expression profiles. miRNAs, a class of non-coding endogenous and short RNAs, are small non-translated modifiers of mRNA expression and translation that regulate gene expression. These groups of miRNAs may show differential levels of expression in benign or malignant processes. Finally, immunohistochemical evaluation can determine the presence or absence at the cellular level of expressed proteins that are associated with benign or malignant tumors. These four categories represent diverse and distinct mechanisms of molecular diagnostic testing, and each has shown the ability to assist in the clinical care of affected patients.5
In this review, we provide information on current methods available for diagnostic and prognostic purposes applied to thyroid nodules. Recent advances in molecular genetics of thyroid nodules can be applied to develop new diagnostic markers for FNA material. A molecular marker assessed on FNA material from a thyroid-nodule biopsy may target one of many cellular components, such as RNA, DNA, and proteins.7 Several specific molecular/immunohistochemical techniques can measure either quantitatively or qualitatively these cellular constituents.
Immunohistochemical theranostic and genomic markers
Histomorphology or cytomorphology remains the cornerstone for the diagnosis of thyroid nodules. Immunohistochemical analysis, by far, is the most commonly used method to complement morphologic assessment.6 The features that help in tumor prognostication are only identified after histologic evaluation. The application of immunohistochemical biomarkers is critical for accurate diagnosis, prognosis, and management of thyroid nodules.8
Hector Battifora mesothelial-1
Hector Battifora mesothelial-1 (HBME-1), an antibody targeting the microvilli of mesothelioma cells, has been detected on differentiated thyroid cancer cells, resulting in its development as a molecular marker for assessing malignancy risk. HBME-1 is measured by immunohistochemistry on formalin-fixed and paraffin-embedded specimens. A preliminary study evaluating HBME-1 on indeterminate cytologic smears have shown moderate test performance, with a sensitivity of 79–94%, specificity 83–94%, positive predictive value (PPV) 84–94%, and negative predictive value (NPV) 80–94%.9
Paired box gene 8
Paired box gene 8 (PAX8) is strongly and diffusely expressed in thyroid follicular epithelium and its associated neoplasms, as well as focal and weak expression in patients with medullary thyroid carcinoma (41–75%).10 It has been demonstrated that PAX8 expression is often retained in the patients with anaplastic thyroid carcinoma and is helpful in cases with limited to no expression of thyroid transcription factor-1 (TTF-1) and cytokeratins, especially those which lack an associated component of well-differentiated thyroid carcinoma.10 It has been also shown that PAX8 immunostaining is helpful to distinguish anaplastic thyroid carcinoma showing squamous differentiation from squamous cell carcinoma of the head and neck, as it is negative in the latter.10
Thyroid transcription factor-2
Thyroid transcription factor-2 (TTF-2) is diffusely and strongly expressed in all well-differentiated of follicular-cell-derived and poorly differentiated carcinomas, and weakly and focally expressed in a few cases of anaplastic carcinoma and in up to 75% of patients with medullary thyroid carcinoma.11
TP53
TP53 is associated with a wide variety of solid-organ malignancies, such as thyroid nodules, and is often associated with late or poorly differentiated disease. In a series of 27 anaplastic carcinoma cases with antecedent papillary thyroid carcinoma (PTC), mutational analysis revealed that BRAFV600E and telomere reverse transcriptase (TERT) mutations are present in 90% and 95% of cases in both components, respectively. Loss of TTF-1 expression and aberrant p53 expression have been exclusively presented in anaplastic carcinoma components in 59% and 63%, respectively, suggesting that these alterations are directly associated with anaplastic transformation. TP53 mutation, which may be detectable as diffuse and intense positivity in immunohistochemistry for p53 protein, is highly prevalent in anaplastic carcinoma but rare in overall PTC.12
Cytokeratin-19 and galectin-3
Cytokeratin-19 (CK19), a cytoskeleton component of epithelial cells, may be up-regulated in well-differentiated thyroid neoplasms. It has been shown that in cytologically in determinate thyroid nodules, most of the nodules ultimately proven to be PTC, stained positive for CK19; however, there was extensive overlap in CK19 expression between benign follicular adenomas and follicular thyroid carcinoma.9
Saleh et al. performed immunohistochemistry staining of galectin-3, HBME-1, Ret oncoprotein (Ret), and CK19, on cell block sections of thyroid FNA biopsy cases that had corresponding surgical resections. They have found that galectin-3 is the best single marker in differentiating benign from malignant thyroid nodules with the highest sensitivity (92.6%) and specificity (77.3%). In addition, it has been demonstrated that the galectin-3 + HBME-1 is the best combination for distinguishing benign from malignant thyroid lesions. Therefore, the use of the galectin-3 + HBME-1 panel is recommended to enhance the diagnostic accuracy of follicular-patterned thyroid lesions on FNA biopsies.13
Diagnostic molecular markers
AFIRMA gene expression classifier
The AFIRMA™ (Veracyte; South San Francisco, California, USA) gene expression classifier (GEC) is a widely employed ‘rule-out’ test, identifying thyroid nodules that are low risk for malignancy so that surgery may be avoided. This test is validated to pre-operatively identify cytologically indeterminate nodules likely to be truly benign or suspicious so that surgery can be avoided.14 Out of these 167 genes, 142 genes represent the most common entities seen in thyroid cancer, the other 25 genes comprising the less common entities seen in the thyroid.15 The effectiveness and value of the Afirma GEC test have been evaluated by several researchers. In a comprehensive prospective study, a total of 4,812 thyroid FNAs from 3,789 patients were evaluated. 16 Out of these samples, 5.52% of the FNA samples were selected for GEC analysis, and the overall sensitivity of the Afirma GEC test was 92% with an NPV of 93% (85% for a lesion suspicious for malignancy, 94% for a follicular neoplasm, and 95% for atypia of undetermined significance/follicular lesion of undetermined significance [AUS/FLUS]).16,17
A 2016 review evaluated the diagnostic accuracy of the GEC over a 6.5-year period by using Quality of Diagnostic Accuracy Studies (QUADAS-2) tool.18 The authors reported specificity, sensitivity, PPV, and NPV of the Afirma GEC in Bethesda III and IV thyroid nodules. Ten of 12 reported studies did not provide a reference standard for GEC-benign thyroid nodules, further highlighting the problem with validating the high NPV reported to date.18 It has been reported that a negative result in the Afirma test has resulted in a considerable decrease in the number of surgeries performed in thyroid lesions classified as categories III and IV; however, a suspicious Afirma result falls below an optimal PPV.19
The detection of medullary thyroid cancer (MTC) is a new addition to GEC testing. A five-gene MTC classifier was developed after using machine learning to select from 283,927 gene transcripts in 22 known MTC cases. It has demonstrated high specificity and sensitivity of 99.8% and 97.9%, respectively, with NPV of 99.8% and PPV of 97.9%.20 However, the targeted next-generation sequencing (NGS) and Afirma GEC tests demonstrated decreases in PPV when noninvasive follicular thyroid neoplasm with papillary-like nuclear features (NIFTYP) were considered nonmalignant.21 It has been recently reported that the Afirma genomic sequencing classifier (GSC) RNAseq panel has high sensitivity and sufficiently high enough specificity to function as both a rule-out and rule-in test.22 The Afirma GEC had lower specificity compared to the targeted NGS in patients with intermediate thyroid nodules and allowed fewer patients to avoid surgery.22 Long-term surveillance is necessary to assess the false-negative rate of these particular molecular tests.22
Genomic sequencing classifier
The novel GSC approach was introduced recently. In a recent prospective study, the sensitivity of the GSC (91%) was similar to that of GEC (89%); however, the specificity was significantly improved (68% for GSC versus 50% for GEC).24 When applied to Bethesda III and IV nodules, GSC offers enhanced performance over GEC by excluding one-third more benign nodules from diagnostic surgery.
MicroRNA classifiers
Microarray platforms are a method of rapidly assessing the expression of hundreds of transcribed RNA sequences at relativly low cost. As computational algorithms are necessary to analyze the expression patterns seen, these techniques may be replaced with NGS platforms; however, it remains a robust form of expression analysis. miRNAs comprise a class of non-coding endogenous and short RNAs (21–23 nucleotides) that regulate gene expression. miRNAs are extremely stable and remain intact in tissues, whether formalin-fixed paraffin-embedded, fresh, or frozen tissues.24 The regulatory role of miRNAs on gene expression is done by directing their target mRNAs for degradation or translational repression.25,26 The role of miRNA expression profiling in cancer diagnosis, prognosis, and response to treatment has been demonstrated. Numerous studies have described the role of miRNAs in the pathogenesis of thyroid cancer.27–30 Rosetta Genomics (Princeton, New Jersey, USA) recently published data for anmiRNA-based assay for diagnosis of Bethesda II–VI thyroid FNA smears.31 A novel assay utilizing miRNA expression in cytology smears was developed. The assay has been developed from a 2,000-miRNA microarray, of which 96 showed differential expressions between malignant and benign thyroid nodules.
The overall sensitivity of the miRNA-based assay was 98% (NPV of 91%) and specificity was 78% (PPV of 59%). Therefore, the need for repeat biopsies or extra passes on initial FNA can be avoided and does not require fresh tissue or special collection and shipment conditions. This test offers a valuable tool for the preoperative classification of thyroid nodules with indeterminate cytology.32 When focusing on Bethesda III and IV samples of ≥1 cm, the sensitivity is 84% (NPV of 92%) and the specificity is 72% (PPV of 43%).32 Over-expression of specific miRNAs has been identified in follicular thyroid versus follicular adenoma and PTC versus normal thyroid tissues.28 Labourier et al. assessed 10-miRNA panel and seven-gene mutational panel on FNA biopsy material from 109 Bethesda III or IV thyroid nodules and demonstrated 74% (95% CI 58–86%) PPV and 94% (95% CI 85–98%) NPV and in a population with a 32% rate of malignancy.33
Affymetrix Exon arrays
Several gene expression microarray profiling studies have been attempted to identify genes that are differentially expressed by benign and malignant thyroid nodules.34,35 The ultimate goal of the microarray profiling assay has been to detect genomic signatures that may be exploited to improve the accuracy of preoperative thyroid cancer diagnosis. The Affymetrix Exon arrays (Affymetrix Inc.; Santa Clara, California, USA) represent an advance in array technology that uses >5.5 million features (previous generation gene-level arrays used approximately 600,000 coding features), and allows for a comprehensive evaluation of every exon in the human genome. Wiseman et al. analyzed 125 thyroid nodules consisting of 65 FNA-biopsy specimens and 60 surgical tissue specimens by using Affymetrix Exon array. The data obtained from a genomic classifier, composed of 249 markers that corresponded to 154 genes, revealed an overall validated accuracy of 90.0% in the 31 patient FNA-biopsy specimens and PPV and NPV of 100% and 85.7%, respectively.36
RosettaGX Reveal
The classification method used for miRNA-based assay, named RosettaGX Reveal™ (Rosetta Genomics; Princeton, New Jersey, USA), combines several linear discriminant analysis steps along with a K-nearest neighbor-based classifier to differentiate between benign and suspicious to malignancy thyroid nodules. Lithwick-Yanai et al. analyzed 156 stained FNA smears, 73 cell blocks of FNAs, and 53 formalin-fixed paraffin-embedded samples of resected tumors, corresponding to 84 unique patients by Agilent custom-designed miRNA microarrays containing over 2,000 miRNA probes. The RosettaGX Reveal showed 98% sensitivity (95% CI 87–100%), 78% specificity (95% CI 69–85%), and 99% NPV (95% CI 94–100%).32
ThyroSeq
ThyroSeq® (CBLPath Inc.; Rye Brook, New York, USA) is a multi-gene test that was initially described by Nikiforova et al. in 2013.31 This test is based on the targeted RNA and DNA NGS analysis of expression levels of 16 genes and mutation in 56 genes (i.e., point mutations and small insertions/deletions in 14 genes and 42 types of gene fusions).37 NGS offers high sensitivity of detection and ability to quantify the proportion of cells carrying a given mutation, meaning that the analysis of mutation hotspots and gene fusions using NGS have been found in ~90% of PTCs and other thyroid cancers.38 Clinical validation of ThyroSeq v2, an improved version of the test, has been reported in two single-institution studies of thyroid nodules with indeterminate cytology and known surgical outcome.39,40
The clinical utility and performance of ThyroSeq v2 have been recently evaluated in several independent studies. Valderrabano et al. assessed 190 thyroid nodules (n=182 patients) with AUS/FLUS and follicular neoplasm/suspicious for follicular neoplasm (FN/SFN) cytology; in this study, the ThyroSeq v2 showed 70% sensitivity, 77% specificity, 42% PPV, and 91% NPV.41 The performance of ThyroSeq v2 was significantly better in FN/SFN than that of AUS/FLUS nodules. A total of five false-negative specimens including two NIFTPs, two PTCs, and one minimally invasive follicular thyroid carcinoma with vascular invasion were reported in their study.41 The authors found that in the thyroid nodules with FN/SFN cytology, a ThyroSeq-positive result increased the risk of malignancy (2.5-fold), with PPV of 53–65%; additionally, ThyroSeq-positive samples (73%) were more frequently resected than that of ThyroSeq-negative samples (48%).41 In another study, ThyroSeq v2 performance was assessed in 148 thyroid nodules with indeterminate cytology. The authors found 95% sensitivity, 60% specificity, 66% PPV, and 94% NPV in their patient population.42 It has been shown that the ThyroSeq had a 100% analytic accuracy in previously positive thyroid tumor samples and cell lines.19
Although ThyroSeq v2 targets 14 genes and 42 gene fusion products with high sensitivity and specificity (more than 90%), independent validation studies have confirmed the high NPV but reported lower PPV.39,40 The development and analytical performance of ThyroSeq v3, an expanded 112-gene version of this molecular test has been recently introduced.43 ThyroSeq v3,a DNA- and RNA-based NGS assay, has been designed by Nikiforova et al. to detect an expanded panel for a variety of genetic alterations/amplification, including 12,135 single-nucleotide variants and deletions/insertions in 112 genes, 120 gene fusions, 19 abnormal gene expression, and 10 copy number alterations in indeterminate thyroid nodules.43 This test uses a genomic classifier to separate benign lesions from malignant lesions. It has been demonstrated that ThyroSeq v3 has improved sensitivity and specificity in the detection of thyroid cancer and improved performance in Hürthle-cell lesions.43
ThyGenX/ThyraMIR
The identification of prevalent oncogenic mutations/translocations in thyroid cancer led to intense focus on the DNA mutations/alterations in thyroid cancer. Early versions of molecular tests, such as the miRInform® (Asuragen; Austin, Texas, USA), utilized a standard panel of four DNA point mutations (BRAF, HRAS, NRAS, KRAS) and three RNA translocation fusion markers (RET/PTC1, RET/PTC3, and PAX8/PPARγ) with high specificity to rule-in malignancy in indeterminate thyroid nodules.19 The more recent ThyGenX® (Interpace Diagnostics, Parsippany, New Jersey, USA) uses NGS to identify >100 genetic alterations across eight genes associated with thyroid malignancy. ThyGenX includes the original genetic alterations/mutations analyzed in the miRInform test as well as testing for PIK3CA mutation, which occurs more frequently in follicular and anaplastic carcinoma.44 BRAF mutation is the most common and specific mutation seen in PTC (incidence of 40–45%); however, RAS and RET/PTC gene mutations are detected in 10–20% of PTCs.5 It is reported that approximately 70% of PTCs carry genetic mutations, either point mutations in the RAS and BRAF genes or present with RET/PTC or TRK rearrangements.45
To improve the false-negative rate of ThyGenX, a multicenter validation of a 10-miRNA gene reflex test, marketed as ThyraMIR® (Interpace Diagnostics, Parsippany, New Jersey, USA), has been performed to improve diagnostic accuracy in thyroid nodules which were screened as 7GP-negative.33 This study have reported PPV 74% and NPV 94%, with calculated PPV >50% and NPV 93–98% for cancer probability of 15–35% when the miRNA classifier was initially trained on 240 surgically resected lesions.33
In another study, the ThyGenX/ThyraMIR pairing was evaluated on a series of 257 FNAs obtained from an academic medical center. The authors used three different testing modalities including ThyraMir, ThyGenX (7GP using a Luminex platform), and an expanded NGS-based mutation panel including 20 genes and 46 fusion transcripts. They found that the addition of miRNA testing increased sensitivity to 94%, compared to 55% with ThyGenX alone.46 Both the NGS-based panel and miRNA testing had optimal sensitivity for thyroid nodules; however, specificity was lower with the expanded genetic panel.46 Truly independent validation of the ThyGenX/ThyraMIR test has yet to be investigated.47
Targeted next-generation sequencing
While molecular markers including existing miRNA and gene expression assays may improve preoperative risk stratification and seem sensitive for detection of malignancy, they lack the ability to provide prognostic information in patients with confirmed malignancy. On the other hand, somatic mutation-based methods provide specific mutation results, which can give important prognostic information.47
Xu et al. performed targeted NGS on eight thyroid tumors, including five encapsulated follicular variants of PTC with invasion (EFVPTC), one infiltrative follicular variant of PTC (IFVPTC), one PTC classical variant, and one Hürthle cell carcinoma. RAS mutations were the main driver in four EFVPTC and one IFVPTC. A classical variant of PTC harbored a BRAFV600E mutation.48 The two tumors, one EFVPTC and one Hürthle cell carcinoma, did not contain RAS or BRAF mutations. In their study, TERT promoter mutations detected at a high frequency (six out of eight tumors), and association between TERT mutation and distant metastasis were demonstrated. A total of 50% of the patients had simultaneous TERT promoter mutations with BRAF, RAS, and BRAFV600E mutations. A RET/PTC rearrangement was not detected in this cohort.48 Co-occurring TERT and BRAFV600E mutations have also been associated with increased risk of tumor recurrence (hazard ratio: 6.74).49 Teng et al. have reported that BRAFV600E was the most common mutation, present in 65% of thyroid nodules, and 94% of the aggressive hobnail variant of PTC.50
Discussion
The aim of this review was to evaluate the sensitivity, specificity, and accuracy, as well as clinical use, of current molecular tests available for diagnostic and prognostic purposes applied to thyroid nodules. It is important to note that, of approximately 600,000 FNAs performed annually in the USA, about 20%, or 100,000 nodules, are identified as indeterminate and subjected to surgery.51 Lobectomy offers final histologic diagnosis, which means most patients are treated unnecessarily.52
Several molecular tests have been proposed to increase diagnostic accuracy in thyroid nodules, including targeted NGS (RA, BRAF, TERT, P53), ThyGenX/ThyraMIR, ThyroSeq (v2 and v3), RosettaGX Reveal, Affymetrix Exon arrays, miRNA classifiers, Galectin-3-ICC (GAL-3-ICC), GEC alone and GEC+BRAF, mutation/fusion (M/F) panel, alone, M/F panel+miRNA GEC, M/F panel by NGS, and TSHR mRNA blood assay. In a comparative review, diagnostic performance, feasibility, and cost of different molecular test-methods for thyroid nodules with indeterminate cytology have been analyzed.53
The ability of a negative diagnostic test to reliably rule out malignancy in a thyroid nodule with indeterminate cytology would be extremely helpful in surgical decision-making, excluding benign lesions that can be selected for follow-up. Additionally, the ability to identify malignancy among indeterminate thyroid nodules would be an important improvement in thyroid practice. The best rule-out and rule-in indicators are the sensitivity and specificity that refer to the proportion of subjects with and without the target condition (malignancy at histology), respectively. Sensitivity gives a positive test result and corresponds to a high NPV and a low false-negative rate, while specificity gives negative test results and corresponds to high PPV and low false-positive rate. Surgical decision based on a good rule-in test may result in clinicians selecting only those with thyroid cancers for surgery, reducing the over-treatments of benign lesions and consequently the social costs. In a recent comparative analysis M/F panel by NGS, GEC alone, GEC + BRAF, and M/F panel + miRNA GEC showed the highest sensitivities (90%, 90%, 89%, and 89%, respectively), the highest NPV (96%, 94%, 92%, and 85%, respectively), and the lowest false-negative rate (10%, 10%, 11%, and 11%, respectively) in the patients with intermediate thyroid nodules.54
Analysis of molecular signature of thyroid nodules has been evolving over the past two decades. The PCR-based screening test for BRAFV600E mutations was initially available with high specificity, PPV, and low sensitivity for detection of thyroid cancer. Sensitivity, specificity, NPV, and PPV of four available molecular tests including Afirma GEC, ThyroSeq v2 and ThyGenX/ThyraMIR and RosettaGX have been systematically investigated to further knowledge about these available molecular tests and understand their advantages and limitations.54
The sensitivity of molecular diagnosis of thyroid nodules was slightly improved by the introduction of ThyGenX, a seven-gene mutation panel, for detecting of the most common genetic alterations including BRAFV600E, NRAS, HRAS, and KRAS mutations present in up to 70% of thyroid cancers and gene fusions RET/PTC1, RET/PTC3, and PAX8–PPARG. ThyGenX has high specificity to rule in malignancy in indeterminate thyroid nodules.19 ThyGenX includes the original genetic alterations/mutations testing for PIK3CA mutation, which occurs frequently in follicular and anaplastic carcinoma, as well as BRAF mutation, which is the most common and specific mutation seen in PTC.32,44 However, the very low NPV of this test precludes its use as a rule-out test.7 To improve the false-negative rate and improve diagnostic accuracy of ThyGenx in thyroid nodules, combination with a multicenter validation of a 10-miRNA gene reflex test, marketed as ThyGenX/ThyraMIR, has been introduced.33 It has been reported that the addition of miRNA testing to ThyGenx (ThyraMIR) increased sensitivity to 94%, compared to 55% with ThyGenX alone.46 However, ThyGenX/ThyraMIR has not been appropriately validated in a large prospective study.55
The RosettaGX Reveal, a miRNA-based assay, has high sensitivity to differentiate between benign and suspicious to malignancy thyroid nodules.32 This test is able to detect 24 miRNA markers with NPV of 92% and PPV of only 43%. Although the potential utility of RosettaGX is in ruling-out of thyroid nodules, the utility of this test for ruling in of thyroid nodules has been limited by its lower specificity.56
ThyroSeq v2 is a test based on the targeted RNA and DNA NGS analysis of expression levels of 16 genes and mutation in 56 genes with high sensitivity and specificity (>90%), high NPV and lower PPV in thyroid nodules of Bethesda IV and Bethesda III cytology.37 Although the clinical utility and performance of ThyroSeq is better in FN/SFN than that of AUS/FLUS nodules, false-negative results in NIFTP, PTC, and minimally invasive follicular thyroid carcinoma have been reported.41 The development and analytical performance of ThyroSeq v3, an expanded 112-gene version of this molecular test has been recently introduced.43 ThyroSeq v3 is another DNA- and RNA-based NGS assay to detect an expanded panel for a variety of genetic alterations/amplification and abnormal gene expression in indeterminate thyroid nodules.43
A meta-analysis of genetic testing methods showed that the sensitivity of the Afirma GEC, ThyroSeq v2 and ThyGenX/ThyraMIR tests for thyroid nodules was consistently close to 90%. While the RosettaGX Reveal showed a sensitivity of 74% when considering the whole cohort, and 100% when non-agreement gold standard cases were excluded. ThyroSeq v2, ThyGenX/ThyraMIR, RosettaGX Reveal, and Afirma GEC have shown specificity of 92%, 85%, 74%, and 52%, respectively. No statistical differences were found for NPV between the four tests, ranging between 92–96%. However, ThyroSeq v2 and ThyGenX/ThyraMIR showed a comparable PPV of 74–78%, which was significantly higher than the PPV than those of both, Afirma GEC and RosettaGX Reveal (37% and 43%, respectively).54
The Afirma GEC is a widely employed rule-out test, identifying thyroid nodules that are low risk for malignancy so that surgery may be avoided. The assay has been designed to analyze the expression profiles of 142 genes based on a proprietary algorithm. The Afirma GEC test is validated to pre-operatively identify cytologically indeterminate thyroid nodules likely to be truly benign or suspicious so that surgery can be avoided.14 Despite of high sensitivity and NPV ranging from 75–100% of this test, the low specificity and PPV ranging only between 14–44%, limiting its use as a rule-in test.16,57,58
In 2015, Nikiforov et al. designed the ThyroSeqNGS panels to improve diagnostic accuracy of thyroid nodules. They designed the first panel, ThyroSeq, to target 12 cancer genes with 284 mutational hot spots. The second panel, ThyroSeq v2, included 56 thyroid-related genes and was introduced to analyze for point mutations and small insertions/deletions in thyroid nodules as well as the expression levels of 16 genes to provide an internal control for an adequacy of cellularity in FNA material.39 The increased number of mutations examined, and the anticipated detection of cancer-specific mutations, should lead to a higher assay sensitivity and be associated with a higher NPV and relatively high specificity and PPV, respectively.40 The high sensitivity and NPV of 96–97% with high specificity associated with PPV of 77–83% of ThyroSeq v2 suggests its potential utility as both a rule-out and a rule-in test.
One of the limitations of this test is the low PPV when the prevalence of cancer in the tested population is low. It has been recently shown that in a population with cancer prevalence of 16%, the NPV of ThyroSeq v2 remained high at 96%, while the PPV was only 22%.59 In contrast to the sensitivity and specificity of any diagnostic test that was affected only by test performance, the PPV and NPV depend on the prevalence of disease in the tested population.
The newest version of ThyroSeq v3 (112-gene test) has been designed to increase sensitivity to rule-out and rule-in by adding recently discovered genetic markers related to thyroid nodules and cancer, including the copy number alteration observed in up to 7% of cancers. Most importantly, ThyroSeq v3 introduces a scoring system to account for molecular signatures typical for both benign adenomas and adenocarcinoma. Another advantage of ThyroSeq v3 is the inclusion of benign and malignant Hürthle-cell lesions.43 Because of the limited inclusion of Hürthle cell carcinoma during test validation and reports of a high false-positive result, the accuracy of molecular testing for Hürthle-cell lesions has been challenged. To address this limitation, ThyroSeq v3 has purposefully trained on a set of surgical specimens enriched in benign and malignant Hürthle-cell neoplasms and was able to distinguish benign from malignant lesions with a sensitivity of 93%. This represents a significant improvement from ThyroSeq v2 and provides reassurance that ThyroSeq v3 can be useful in the evaluation of Hürthle-cell neoplasms.43
Although molecular markers can improve preoperative risk stratification and seem sensitive for detection of malignancy, they lack the ability to provide prognostic information in patients with confirmed malignancy. Unlike the aforementioned molecular tests, targeted NGS (RAS, BRAF, TERT, P53) or somatic mutation-based methods provide specific mutation results, which can give important prognostic information.47 However, the performance of all of the molecular tests is also unknown for NIFTP, as it has previously been categorized as either benign or malignant noninvasive EFVPTC. Moreover, one of the potential limitations of the routine utilization of molecular diagnostics of thyroid nodules is their relatively high cost. Lee et al. concluded that the Afirma assay was not associated with beneficial cost-effectiveness in the Canadian healthcare system, but was in the US system.60
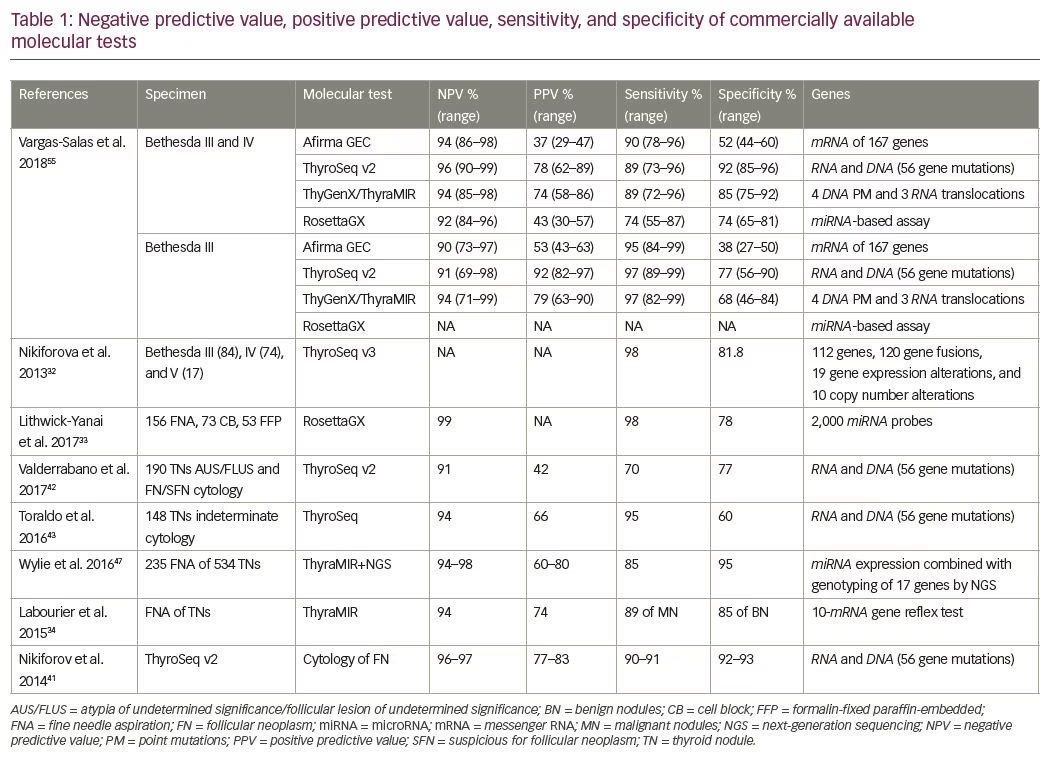
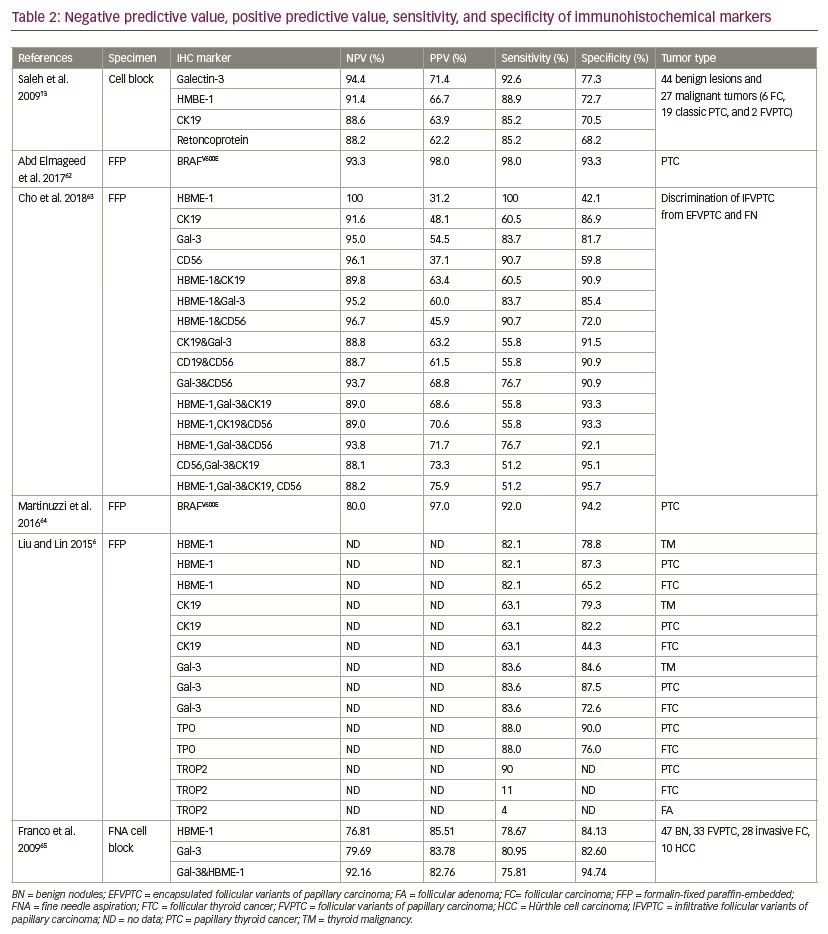
It has been suggested that interpretation of the molecular results of indeterminate thyroid nodules is affected by cancer prevalence at a given institution. A high cancer prevalence increases the PPV, whereas low cancer prevalence increases NPV of the test. Finally, proper clinical judgment is best based on presence of comorbidities, patient age, ultrasonographic features of the thyroid nodules, and identification of molecular signature to select candidates for active surveillance or surgery.57 The sensitivity, specificity, PPV, and NPV of each molecular method are presented in Tables 1 and 2.6,13,31–33,40–42,46,54,61–64
Conclusions
There is no single means of evaluation that will effectively diagnose an indeterminate nodule as benign or malignant with high accuracy. This accuracy has been recently improved through the application of specific molecular markers or molecular profiles of thyroid nodules correlated with benign or malignant disease, and has reduced unnecessary surgery while enhancing understanding of individualized prognosis. Unfortunately, the features that help in thyroid tumor prognostication are only identified after histologic evaluation. Molecular markers may improve preoperative risk stratification. Although existing miRNA and gene expression assays have been shown to be sensitive for detection of malignancy, they lack the ability to provide prognostic information in patients with confirmed malignancy. Somatic mutation-based testing and immunohistochemical markers, however, provide specific mutation results, which can provide important prognostic information. Currently, four commercially available molecular tests are useful for ruling out malignancy in indeterminate thyroid nodules; however, long-term follow-up studies are needed to confirm that test-negative thyroid nodules are truly benign. Additionally, the predictive abilities of the test are also variable based upon regional differentiated thyroid cancer prevalence and genetic signatures. Despite these limitations, molecular markers used in the correct setting can help clinicians and patients to decide on management strategies in indeterminate thyroid nodules.